Training of DnCNN for Denoising#
This example demonstrates the training and application of the DnCNN model from [58] to denoise images that have been corrupted plot.config_notebook_plotting() with additive Gaussian noise.
[1]:
import os
from time import time
import numpy as np
import jax
from mpl_toolkits.axes_grid1 import make_axes_locatable
from scico import flax as sflax
from scico import metric, plot
from scico.flax.examples import load_image_data
Prepare parallel processing. Set an arbitrary processor count (only applies if GPU is not available).
[2]:
os.environ["XLA_FLAGS"] = "--xla_force_host_platform_device_count=8"
platform = jax.lib.xla_bridge.get_backend().platform
print("Platform: ", platform)
Platform: gpu
Read data from cache or generate if not available.
[3]:
size = 40 # patch size
train_nimg = 400 # number of training images
test_nimg = 16 # number of testing images
nimg = train_nimg + test_nimg
gray = True # use gray scale images
data_mode = "dn" # Denoising problem
noise_level = 0.1 # Standard deviation of noise
noise_range = False # Use fixed noise level
stride = 23 # Stride to sample multiple patches from each image
train_ds, test_ds = load_image_data(
train_nimg,
test_nimg,
size,
gray,
data_mode,
verbose=True,
noise_level=noise_level,
noise_range=noise_range,
stride=stride,
)
Data read from path : ~/.cache/scico/examples/data
Set --training-- : Size: 104000
Set --testing -- : Size: 16640
Data range -- images -- : Min: 0.00, Max: 1.00
Data range -- labels -- : Min: 0.00, Max: 1.00
NOTE: If blur kernel or noise parameter are changed, the cache must be manually deleted to ensure that the training data is regenerated with these new parameters.
Define configuration dictionary for model and training loop.
Parameters have been selected for demonstration purposes and relatively short training. The depth of the model has been reduced to 6, instead of the 17 of the original model. The suggested settings can be found in the original paper.
[4]:
# model configuration
model_conf = {
"depth": 6,
"num_filters": 64,
}
# training configuration
train_conf: sflax.ConfigDict = {
"seed": 0,
"opt_type": "ADAM",
"batch_size": 128,
"num_epochs": 50,
"base_learning_rate": 1e-3,
"warmup_epochs": 0,
"log_every_steps": 5000,
"log": True,
"checkpointing": True,
}
Construct DnCNN model.
[5]:
channels = train_ds["image"].shape[-1]
model = sflax.DnCNNNet(
depth=model_conf["depth"],
channels=channels,
num_filters=model_conf["num_filters"],
)
Run training loop.
[6]:
workdir = os.path.join(os.path.expanduser("~"), ".cache", "scico", "examples", "dncnn_out")
train_conf["workdir"] = workdir
print(f"{'JAX process: '}{jax.process_index()}{' / '}{jax.process_count()}")
print(f"{'JAX local devices: '}{jax.local_devices()}")
trainer = sflax.BasicFlaxTrainer(
train_conf,
model,
train_ds,
test_ds,
)
start_time = time()
modvar, stats_object = trainer.train()
time_train = time() - start_time
JAX process: 0 / 1
JAX local devices: [cuda(id=0), cuda(id=1), cuda(id=2), cuda(id=3), cuda(id=4), cuda(id=5), cuda(id=6), cuda(id=7)]
Channels: 1, training signals: 104000, testing signals: 16640, signal size: 40
+---------------------------------+----------------+--------+-----------+--------+
| Name | Shape | Size | Mean | Std |
+---------------------------------+----------------+--------+-----------+--------+
| ConvBNBlock_0/BatchNorm_0/bias | (64,) | 64 | 0.0 | 0.0 |
| ConvBNBlock_0/BatchNorm_0/scale | (64,) | 64 | 1.0 | 0.0 |
| ConvBNBlock_0/Conv_0/kernel | (3, 3, 64, 64) | 36,864 | -0.000522 | 0.0589 |
| ConvBNBlock_1/BatchNorm_0/bias | (64,) | 64 | 0.0 | 0.0 |
| ConvBNBlock_1/BatchNorm_0/scale | (64,) | 64 | 1.0 | 0.0 |
| ConvBNBlock_1/Conv_0/kernel | (3, 3, 64, 64) | 36,864 | 0.000178 | 0.0589 |
| ConvBNBlock_2/BatchNorm_0/bias | (64,) | 64 | 0.0 | 0.0 |
| ConvBNBlock_2/BatchNorm_0/scale | (64,) | 64 | 1.0 | 0.0 |
| ConvBNBlock_2/Conv_0/kernel | (3, 3, 64, 64) | 36,864 | 5.46e-05 | 0.0588 |
| ConvBNBlock_3/BatchNorm_0/bias | (64,) | 64 | 0.0 | 0.0 |
| ConvBNBlock_3/BatchNorm_0/scale | (64,) | 64 | 1.0 | 0.0 |
| ConvBNBlock_3/Conv_0/kernel | (3, 3, 64, 64) | 36,864 | -9.56e-05 | 0.0592 |
| conv_end/kernel | (3, 3, 64, 1) | 576 | -0.00121 | 0.0605 |
| conv_start/kernel | (3, 3, 1, 64) | 576 | 0.0155 | 0.457 |
+---------------------------------+----------------+--------+-----------+--------+
Total: 149,120
+--------------------------------+-------+------+------+-----+
| Name | Shape | Size | Mean | Std |
+--------------------------------+-------+------+------+-----+
| ConvBNBlock_0/BatchNorm_0/mean | (64,) | 64 | 0.0 | 0.0 |
| ConvBNBlock_0/BatchNorm_0/var | (64,) | 64 | 1.0 | 0.0 |
| ConvBNBlock_1/BatchNorm_0/mean | (64,) | 64 | 0.0 | 0.0 |
| ConvBNBlock_1/BatchNorm_0/var | (64,) | 64 | 1.0 | 0.0 |
| ConvBNBlock_2/BatchNorm_0/mean | (64,) | 64 | 0.0 | 0.0 |
| ConvBNBlock_2/BatchNorm_0/var | (64,) | 64 | 1.0 | 0.0 |
| ConvBNBlock_3/BatchNorm_0/mean | (64,) | 64 | 0.0 | 0.0 |
| ConvBNBlock_3/BatchNorm_0/var | (64,) | 64 | 1.0 | 0.0 |
+--------------------------------+-------+------+------+-----+
Total: 512
Initial compilation, this might take some minutes...
Initial compilation completed.
Epoch Time Train_LR Train_Loss Train_SNR Eval_Loss Eval_SNR
---------------------------------------------------------------------
6 4.83e+01 0.001000 0.001943 13.21 0.001000 14.17
12 9.17e+01 0.001000 0.000936 14.34 0.001007 14.14
18 1.35e+02 0.001000 0.000874 14.64 0.000933 14.47
24 1.79e+02 0.001000 0.000842 14.80 0.000892 14.68
30 2.21e+02 0.001000 0.000820 14.91 0.000865 14.81
36 2.64e+02 0.001000 0.000808 14.98 0.000842 14.93
43 3.07e+02 0.001000 0.000801 15.02 0.001132 13.72
49 3.49e+02 0.001000 0.000795 15.05 0.000960 14.40
Evaluate on testing data.
[7]:
test_patches = 720
start_time = time()
fmap = sflax.FlaxMap(model, modvar)
output = fmap(test_ds["image"][:test_patches])
time_eval = time() - start_time
output = np.clip(output, a_min=0, a_max=1.0)
Evaluate trained model in terms of reconstruction time and data fidelity.
[8]:
snr_eval = metric.snr(test_ds["label"][:test_patches], output)
psnr_eval = metric.psnr(test_ds["label"][:test_patches], output)
print(
f"{'DnCNNNet training':18s}{'epochs:':2s}{train_conf['num_epochs']:>5d}"
f"{'':21s}{'time[s]:':10s}{time_train:>7.2f}"
)
print(
f"{'DnCNNNet testing':18s}{'SNR:':5s}{snr_eval:>5.2f}{' dB'}{'':3s}"
f"{'PSNR:':6s}{psnr_eval:>5.2f}{' dB'}{'':3s}{'time[s]:':10s}{time_eval:>7.2f}"
)
DnCNNNet training epochs: 50 time[s]: 355.81
DnCNNNet testing SNR: 15.17 dB PSNR: 27.80 dB time[s]: 4.67
Plot comparison. Note that patches have small sizes, thus, plots may correspond to unidentifiable fragments.
[9]:
np.random.seed(123)
indx = np.random.randint(0, high=test_patches)
fig, ax = plot.subplots(nrows=1, ncols=3, figsize=(15, 5))
plot.imview(test_ds["label"][indx, ..., 0], title="Ground truth", cbar=None, fig=fig, ax=ax[0])
plot.imview(
test_ds["image"][indx, ..., 0],
title="Noisy: \nSNR: %.2f (dB), PSNR: %.2f"
% (
metric.snr(test_ds["label"][indx, ..., 0], test_ds["image"][indx, ..., 0]),
metric.psnr(test_ds["label"][indx, ..., 0], test_ds["image"][indx, ..., 0]),
),
cbar=None,
fig=fig,
ax=ax[1],
)
plot.imview(
output[indx, ..., 0],
title="DnCNNNet Reconstruction\nSNR: %.2f (dB), PSNR: %.2f"
% (
metric.snr(test_ds["label"][indx, ..., 0], output[indx, ..., 0]),
metric.psnr(test_ds["label"][indx, ..., 0], output[indx, ..., 0]),
),
fig=fig,
ax=ax[2],
)
divider = make_axes_locatable(ax[2])
cax = divider.append_axes("right", size="5%", pad=0.2)
fig.colorbar(ax[2].get_images()[0], cax=cax, label="arbitrary units")
fig.show()

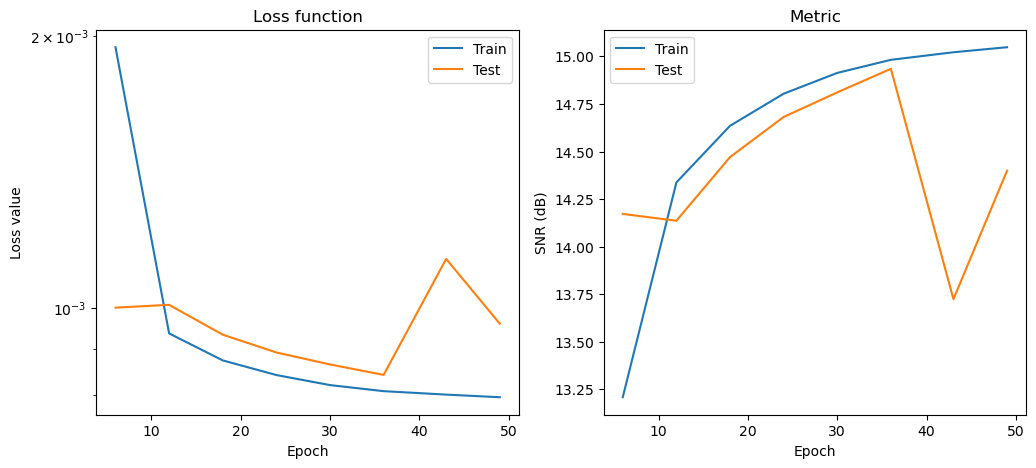
Plot convergence statistics. Statistics are generated only if a training cycle was done (i.e. if not reading final epoch results from checkpoint).
[10]:
if stats_object is not None and len(stats_object.iterations) > 0:
hist = stats_object.history(transpose=True)
fig, ax = plot.subplots(nrows=1, ncols=2, figsize=(12, 5))
plot.plot(
np.vstack((hist.Train_Loss, hist.Eval_Loss)).T,
x=hist.Epoch,
ptyp="semilogy",
title="Loss function",
xlbl="Epoch",
ylbl="Loss value",
lgnd=("Train", "Test"),
fig=fig,
ax=ax[0],
)
plot.plot(
np.vstack((hist.Train_SNR, hist.Eval_SNR)).T,
x=hist.Epoch,
title="Metric",
xlbl="Epoch",
ylbl="SNR (dB)",
lgnd=("Train", "Test"),
fig=fig,
ax=ax[1],
)
fig.show()